The Volmer Group is broadly interested in fundamental and applied aspects of biological mass spectrometry, with current focus on clinical mass spectrometry, metabolomics, high-resolution mass spectrometry of complex natural organic matter and instrument development. Please browse through our publications for an overview of general research interests. Below are highlights of our current work.

Analytical approaches to quantify low-abundant dynamic metabolites in the vitamin D metabolic cascade
The goal of our vitamin D research is the development of novel mass spectrometric techniques to provide detailed insight into the vitamin D metabolome. Vitamin D comprises a group of secosteroidal compounds, of which vitamin D3 is the most important bioactive variant. Vitamin D plays a crucial role in bone health but has also been linked to many other diseases such as cancer and chronic liver diseases. Current clinical assays often lack specificity and accuracy, therefore MS-based techniques are preferred today. We have established several novel MS-based for vitamin D metabolites. We have also developed powerful chemical derivatization tools that increase both sensitivity and specificity of analysis, thus permitting quantitative measurement of multiple vitamin D metabolites from sera of patients. In addition, we have implemented a novel calibration technique for vitamin D from dried blood spots. Currently, we develop novel MS techniques that allow much deeper access into the Vitamin D metabolome than presently possible, to capture very low abundant, dynamic metabolites of vitamin D. We are particularly interested in the parallel epimerization pathway of vitamin D as well as free vitamin D and its protein-bound species. Finally, we are systematically examining the spectrum and dynamic changes of a large number of vitamin D metabolites in sera of patients, to evaluate the diagnostic potential of these fingerprints.
This work is funded by the German Research Foundation (DFG VO 1355/5-1 and VO 1355/5-2).
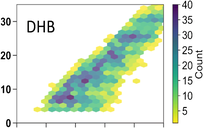
Novel MS-based analytical approaches for shotgun lignomics and quantification of lignin
The goal of our lignin research is the development of new MS techniques for lignomics research. The lignome in this context is the full complement of lignin-related compounds in a given lignin sample; that is, all lignin monomers, oligomers and polymers. Lignin is a major component of woody plants and second most abundant natural polymer, but difficult to degrade chemically. Depolymerization is extremely important, however, for recovery of value chemicals and biofuels from lignin wastes. We have developed novel FTICR-MS techniques that showed that electrochemical degradation of lignin yielded >5000 breakdown products. We have also developed powerful 2D visualization tools that provide detailed information on the transformation and exact degradation mechanisms as well as on the identity of the cleavage products. Despite these powerful approaches, MS is not capable to characterize full lignomes and also exhibits shortcomings for analysis of very low abundant species as well as isomers and isobars. We currently develop novel shotgun MS methods that dig much deeper into the lignome and also yield quantitative information on lignin subunits. Lignin sequencing is difficult because of the heterogenous nature, the crosslinking and the lack of linear repeating patterns. We are conducting comprehensive experiments using MS/MS, including electron-activated dissociation (ExD), to obtain detailed information on the different linkages of lignin subunits. Our strategy also comprises ion mobility spectrometry, to separate the numerous isomers and isobars prior to MS. Finally, by developing enhanced data mining strategies for the complex mass spectral raw data, we obtain new quantitative tools that permit improved quantitative determination of lignin subunits.
This work is funded by the German Research Foundation (DFG VO 1355/4-1 and VO1355/4-3).

Multimodal clinical mass spectrometry to target treatment resistance
– MSTARS Berlin Consortium –
After significant progress of genomics-driven approaches during the past decade, today’s frontier to precision medicine is to gain profound understanding of human diseases at the level of proteins and metabolites. This knowledge holds the promise to improve patient management decisions by providing predictive biomarkers, explanations for treatment failures in individual patients, and by informing future drug development and clinical trial design. We will build an integrated multimodal mass spectrometry core to target treatment resistance. Our research core covers a broad range of complementary proteomic, metabolomic and imaging-based technologies and is complemented by dedicated computational groups for data management, modeling, and machine learning. We will follow two complementary research strategies: (1) a mechanistic strategy that leads from perturbation experiments in preclinical models to predictive biomarkers and (2) a signature-driven strategy that seeks to deduce predictive signatures directly from precisely analyzed patient samples in large informative cohorts. While our research concept is generic, we will focus on cancer and inflammatory disease, with patient cohorts selected for medical need and availability of high-quality clinical samples. Our central use case is head and neck cancer, a devastating disease for which predictive assays are urgently needed. A worldwide unique collection of clinical samples and patient-derived preclinical models available to us will provide the ideal basis for the consolidation of our research core. The multimodal data obtained will be systematically integrated and modeled to predict clinical outcome, to guide decision making, and to design predictive assays for routine application. Our goal is to bundle locally available expertise in mass spectrometry to build a sustainable platform for the benefit of future patients.
This work is funded by the German Federal Ministry of Education and Research (BMBF 031L0220C).
BUA proteomics and metabolomics core computational Link lab (joint HU, TU and Charité facility)
The MS Coputational Link Lab serves as an advanced central computational resource for omics-based mass spectrometry research in Berlin. This resource bridges the rapidly widening gap between the possibilities of ‘Big Data’ interpretation and the production of such data by mass spectrometry. Importantly, no single mass spectrometry lab, while excellent in designing experiments as well as recording and processing data, can cope with the accelerating number of new developments that help in its biomedical interpretation. The Link lab acts as a single point of contact for BUA investigators seeking support to pursue mass spectrometry-based omics research and for outside collaborators wishing to engage with BUA scientists in this area.
This projected is funded by the Berlin University Alliance (BUA 501_LinkLab)
